 (Left to right): BBI's Jimmy Bennett, Evan Eichler, and Andrew Stergachis
(Left to right): BBI's Jimmy Bennett, Evan Eichler, and Andrew Stergachis
Researchers at the University of Washington School of Medicine and Seattle Children’s Research Institute (SCRI) in Seattle have been awarded a $12.5 million grant from the U.S. National Institutes of Health (NIH) to study variations in the genomes of normal cells that accumulate as we grow, develop and age.
The UW effort will be led by Dr. Andrew B. Stergachis, assistant professor of medicine; Dr. Jimmy Bennett, associate professor of pediatrics and principal investigator in the Center for Developmental Biology and Regenerative Medicine at the Seattle Children’s Research Institute; and Evan Eichler, professor of genome sciences. The grant is part of a $140-million, five-year effort led by NIH to characterize the differences between the genomes of somatic cells.
Somatic cells arise from the fertilized egg, or zygote, and compose most of the body’s tissues and organs. As these cells grow, divide and transform into different cell types, their genomes accumulate changes in the DNA sequences. As a result, the genomes of different somatic cell lines differ not only from the genome of the fertilized egg from which they arose, but also from the genomes of each other, as well.
This variation is called somatic mosaicism.
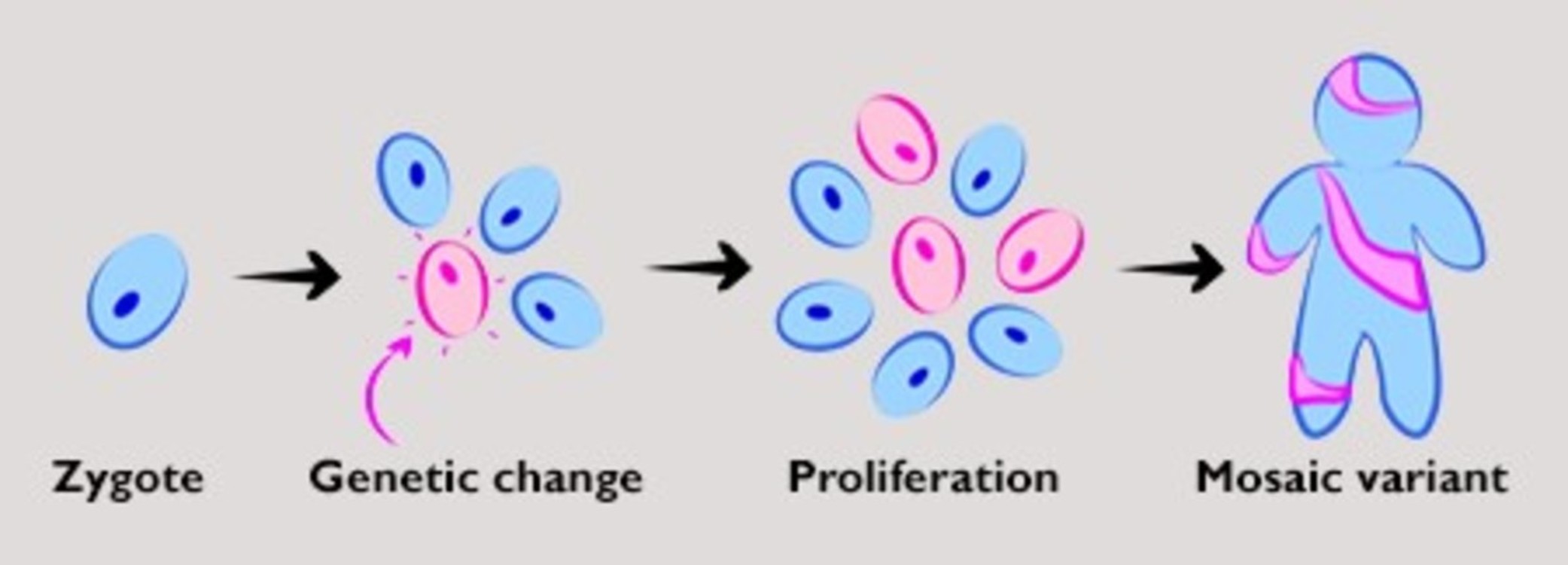 Mutations in cells derived from the zygote can create variant cell lines that appear in some tissues but not others, resulting in what is called mosaic variation. Illustration by Dana Jensen, Bennett Lab
Mutations in cells derived from the zygote can create variant cell lines that appear in some tissues but not others, resulting in what is called mosaic variation. Illustration by Dana Jensen, Bennett Lab
“We tend to think that each of us has a single genome, but the genomes of our cells differ because copying errors made during cell division,” explained Bennett, who studies how post-zygotic mutations affect human development and birth defects in children. “We want to determine the baseline rate of changes in the genomes across multiple human tissues and how they affect biology and health.”
The goal of the new NIH program, called the Somatic Mosaicism across Human Tissues (SMaHT) Network, is to begin to systematically catalog the genomic differences across a wide variety of normal human tissues.
The UW group is one of 22 that have been awarded funding to establish the network. The network will start by sequencing and characterizing somatic genetic variation in tissues from 150 deceased human donors who represent diverse ancestries and represent different life stages.
The UW-SCRI effort will analyze genomic variation across the entire genomes of cells from 10 different tissues from 50 donors.
Determining the complete sequences of these genomes is essential because many of the variants known to cause disease occur in genomic regions that are technically very difficult to sequence, Stergachis said.
“We know these regions play an important role in the development of core features of human biology as well as common human diseases, but we have a poor handle on what is going on in them. Professor Eichler’s team is a world leader in mapping and evaluating these structurally complex variant regions," he said.
In addition to complete sequencing of the genomes from end to end, the UW researchers will characterize molecular modifications to genes and chromosomes that do not affect their DNA sequence but still alter their expression. Such “epigenetic” modifications, which can determine whether a gene is active or inactive, can be passed down from cell to cell, affecting their growth, development and function — for generations.
“Our cells read their genomes with proteins that interact with the DNA,” Sterghacis explained. “In my lab, we have developed a technique called Fiber-seq that tells us where these proteins are bound along the genome. This helps tell us how somatic genetic changes alter where our proteins bind and how they interpret the genome in that segment.”
“With all the other members of the SMaHT network, we hope to produce not only the most complete catalog of somatic variation, but also to learn how these variations impact the ability of cells to function,” he added.
The project is funded by the NIH’s Common Fund (1UM1DA058220-01), which supports research challenges that cannot be addressed by a single NIH institute or center.


